FCU_cardiacAP_composite
About this Functional Cell Unit (FCU)
This is a workspace containing bond graph models of ion channels and transporters which were identified to play important roles in the generation and maintenance of an action potential within a cardiac myocyte. Simply put, an ion channel transports ion(s) from one cellular compartment to another.
One can select components from what is available and build their own composite CellML file.
An existing CellML file is provided, which contains the channels used by Kernik et. al., except for the 'funny' current channel and the 'transient outward' channel.
The list of channels in this workspace is subject to change as literature demands it.
For a given cell model:
- INPUTS:
- Stimulus current (Istim)
- OUTPUTS:
- Cytosolic and extracellular ionic concentrations (Na, K, Ca)
- Membrane potential (Vm)
- Currents through each channel (I)
- CHANNELS AND TRANSPORTERS:
Abbreviation Name NaB Background sodium channel fast-Na Fast sodium channel K1 Inward rectifying potassium channel Kr Rapid delayed rectifying potassium channel Ks Slow delayed rectifying potassium channel CaB Background calcium channel Caleak Leakage calcium channel LCC L-type calcium transporter TCC T-type calcium transporter pCa Sarcolemmal calcium pump NaK Sodium-potassium pump NCX Sodium-calcium exchanger RyR Ryanodine receptor SERCA Sarcoplasmic-endoplasmic reticulum calcium pump CMDN Calmodulin buffer for calcium TRPN Troponin buffer for calcium
Model status
For individual channels, their current CellML implementation run in OpenCOR. Composite models do not run due to solver failure, including the model reproducing Kernik et. al. An exported MATLAB script is provided as an alternative.
Model overview
The components of this workspace are based on existing kinetic models taken from Kernik et. al. (2019) or from Luo Rudy (1994), where the mathematics are translated into the bond-graph formalism. This describes the model in energetic terms and forces adherence to the laws of thermodynamics.
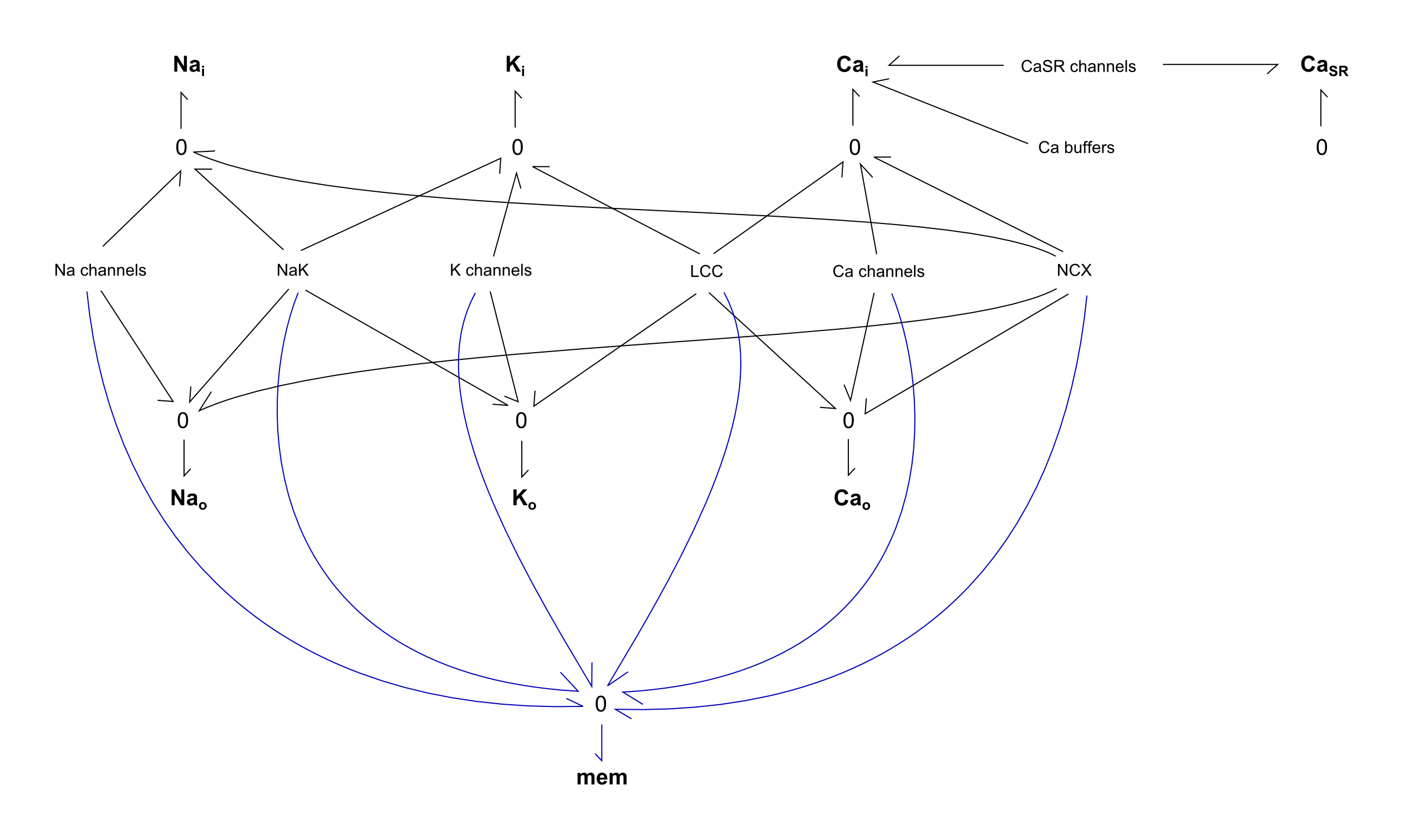
Fig. 1. Bond-graph formulation of the cardiac action potential FCU, following the Kernik model. Na channels consist of fast-Na, NaB; K channels consist of K1, Kr, Ks; Ca channels consist of CaB, TCC, pCa (not voltage dependent); CaSR channels consist of RyR, SERCA, Caleak; Ca buffers consist of CMDN, TRPN. Subscripts i, o and SR refer to the cytosolic, extracellular and sarcoplasmic compartments, respectively. Adapted from Pan et. al..
For an individual channel:
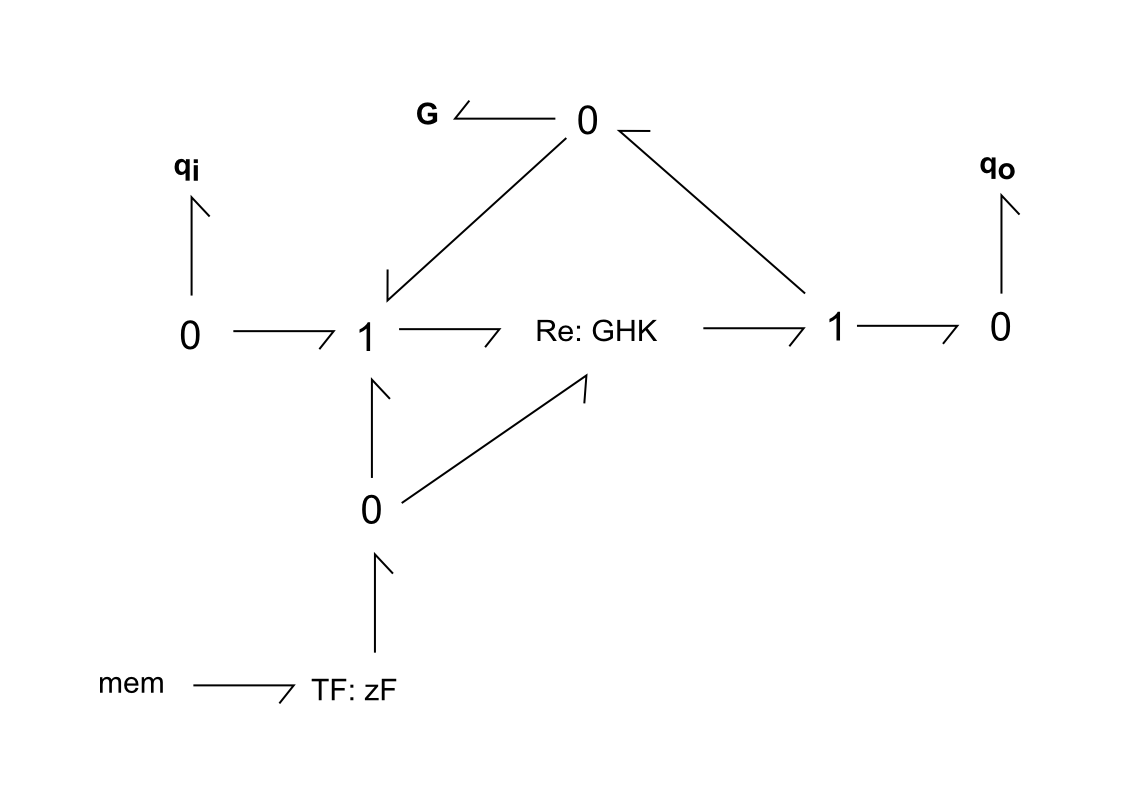
Fig. 2. Bond-graph formulation of an individual gated channel, for the transport of ion q from compartment i to o. G represents the gate, for gated channels. Membrane voltage mem modulates the GHK conductance of the channel and the gate. Adapted from Pan et. al.
For the above bond-graphs, a '0' node refers to a junction where all chemical potentials are the same. A '1' node refers to all fluxes being the same going in and out of the junction.
The following is a list of all available channels of this workspace. Unless otherwise stated, a source index of '1' indicates the model was based on Kernik; a source of '2' on the Luo-Rudy 1994 model; '3' indicates that the bond-graph model was made by Pan et. al. (and mostly based on Luo Rudy). For cases '2' and '3', models have been reparameterised with Kernik cell dimensions.
| Abbreviation | Name | Kinetic source |
|---|---|---|
| CaB | Background calcium channel | 2 |
| Caleak | Leakage calcium channel | 1 |
| CMDN | Calmodulin buffer for calcium | 3 |
| fast-Na | Fast sodium channel | 3 |
| K | Time-dependent potassium channel | 3 |
| K1 | Inward rectifying potassium channel | 3 |
| KATP | ATP-sensitive potassium channel | Clancy and Rudy |
| Kr | Rapid delayed rectifying potassium channel | 1 |
| Ks | Slow delayed rectifying potassium channel | 1 |
| LCC | L-type calcium transporter | 3 |
| NaB | Background sodium channel | 2 |
| NaK | Sodium-potassium pump | 3 |
| NCX | Sodium-calcium exchanger | 3 |
| NS | Non-specific calcium activated channel | 2 |
| pCa | Sarcolemmal calcium pump | 1 |
| RyR | Ryanodine receptor | Stern et. al. |
| SERCA | Sarcoplasmic-endoplasmic reticulum calcium pump | 3 |
| TCC | T-type calcium transporter | 1 |
| TRPN | Troponin buffer for calcium | 3 |
Parameter finding
A description of the process to find bond-graph parameters is shown in the folder parameter_finder, which relies on the:
- stoichiometry of system
- kinetic constants for forward/reverse reactions
- If not already, all reactions are made reversible by assigning a small value to the reverse direction.
- linear algebra script, which outputs the bond-graph parameters K and κ.
Here, this solve process is performed in Python. For a given composite model, the user must specify which components to include through the variable subsystem_names. The stoichiometry for the whole system is automatically found.
Notes on creating a composite model
- Step 3 (above) must be run each time a change is made to the composite model (e.g. addition or removal of a component).
- Kinetic parameters fitted to a kinetic model must be found anew for a cell with different volumes and membrane capacitance values for each individual channel. This would apply to the GHK fits to a linear (ohmic) current, and to finding transition parameters for gating subcomponents of a gated channel. Following this, the bond-graph parameters are then recalculated.
- New channels must have species names consistent with names already present in the model.
For further reading into the derivation of this process, please see Pan et. al..
